| Home | Overview of gut regions | Anatomy | Histology | Transgene expression mapping | Gene expression |
| Search expression data by gene: |
| Gene name | Adh | ||||||||||||||||||||||||||||||||||||
| Flybase description | The gene Alcohol dehydrogenase is referred to in FlyBase by the symbol Dmel\Adh (CG3481, FBgn0000055). | ||||||||||||||||||||||||||||||||||||
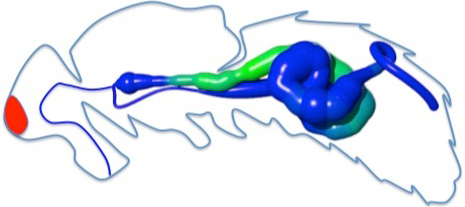
| Expression data along the gut |
|
||||||||||||||||||||||||||||||||||||
| Intestinal gene expression in different physiological conditions |
Ecc15: flies orally infected with Erwinia carotovora carotovora 15. Pe: flies orally infected with Pseudomonas entomophila. Pe gacA: flies orally infecte with Pseudomonas entomophila gacA. For methods and description, see Buchon et al. 2009, Cell Host Microbe, and Chakrabarti et al. 2012, Cell Host Microbe. |
||||||||||||||||||||||||||||||||||||
| Gene details (from Flybase) | It is a protein_coding_gene from Drosophila melanogaster. There is experimental evidence that it has the molecular function: protein homodimerization activity; alcohol dehydrogenase (NAD) activity; acetaldehyde dehydrogenase (acetylating) activity. There is experimental evidence that it is involved in the biological process: alcohol catabolic process; alcohol metabolic process; ethanol oxidation; behavioral response to ethanol; acetaldehyde metabolic process; ethanol metabolic process. 587 alleles are reported. No phenotypic data is available. It has 5 annotated transcripts and 5 annotated polypeptides. Protein features are: Alcohol dehydrogenase, Drosophila-type; Alcohol dehydrogenase, insect-type; NAD(P)-binding domain; Short-chain dehydrogenase/reductase SDR; Short-chain dehydrogenase/reductase, conserved site. Summary of modENCODE Temporal Expression Profile: Temporal profile ranges from a peak of extremely high expression to a trough of low expression. Peak expression observed at stages throughout the larval period, in adult male stages. This gene is annotated by FlyBase as a dicistronic gene, meaning that some or all of its transcripts encode two or more polypeptide-coding open reading frames (ORFs) , with each ORF assigned to a different gene. The distribution of RNA-Seq coverage data amongst the different encoded genes cannot be determined. . |