| Home | Overview of gut regions | Anatomy | Histology | Transgene expression mapping | Gene expression |
| Search expression data by gene: |
| Gene name | Gad1 | ||||||||||||||||||||||||||||||||||||
| Flybase description | The gene Glutamic acid decarboxylase 1 is referred to in FlyBase by the symbol Dmel\Gad1 (CG14994, FBgn0004516). | ||||||||||||||||||||||||||||||||||||
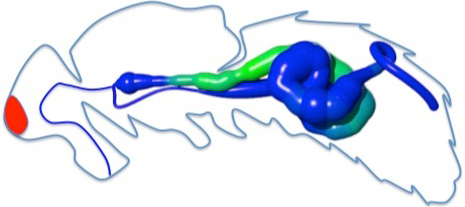
| Expression data along the gut |
|
||||||||||||||||||||||||||||||||||||
| Intestinal gene expression in different physiological conditions |
Ecc15: flies orally infected with Erwinia carotovora carotovora 15. Pe: flies orally infected with Pseudomonas entomophila. Pe gacA: flies orally infecte with Pseudomonas entomophila gacA. For methods and description, see Buchon et al. 2009, Cell Host Microbe, and Chakrabarti et al. 2012, Cell Host Microbe. |
||||||||||||||||||||||||||||||||||||
| Gene details (from Flybase) | It is a protein_coding_gene from Drosophila melanogaster. There is experimental evidence that it has the molecular function: glutamate decarboxylase activity. There is experimental evidence that it is involved in the biological process: response to mechanical stimulus; synapse assembly; gamma-aminobutyric acid biosynthetic process; neuromuscular junction development; neurotransmitter receptor metabolic process; glutamate catabolic process; larval locomotory behavior. 19 alleles are reported. The phenotypes of these alleles are annotated with: bouton; neuromuscular junction. It has 3 annotated transcripts and 3 annotated polypeptides. Protein features are: Pyridoxal phosphate-dependent decarboxylase; Pyridoxal phosphate-dependent transferase, major domain; Pyridoxal phosphate-dependent transferase, major region, subdomain 1; Pyridoxal phosphate-dependent transferase, major region, subdomain 2; Pyridoxal-phosphate binding site. Summary of modENCODE Temporal Expression Profile: Temporal profile ranges from a peak of high expression to a trough of extremely low expression. Peak expression observed within 18-24 hour embryonic stages, during late pupal stages. . |