| Home | Overview of gut regions | Anatomy | Histology | Transgene expression mapping | Gene expression |
| Search expression data by gene: |
| Gene name | Tim10 | ||||||||||||||||||||||||||||||||||||
| Flybase description | The gene Translocase of inner membrane 10 is referred to in FlyBase by the symbol Dmel\Tim10 (CG9878, FBgn0027360). | ||||||||||||||||||||||||||||||||||||
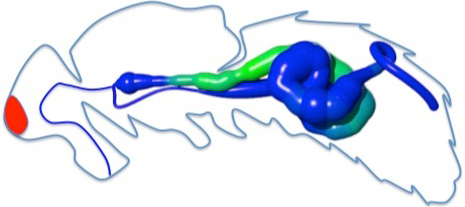
| Expression data along the gut |
|
||||||||||||||||||||||||||||||||||||
| Intestinal gene expression in different physiological conditions |
Ecc15: flies orally infected with Erwinia carotovora carotovora 15. Pe: flies orally infected with Pseudomonas entomophila. Pe gacA: flies orally infecte with Pseudomonas entomophila gacA. For methods and description, see Buchon et al. 2009, Cell Host Microbe, and Chakrabarti et al. 2012, Cell Host Microbe. |
||||||||||||||||||||||||||||||||||||
| Gene details (from Flybase) | It is a protein_coding_gene from Drosophila melanogaster. Based on sequence similarity, it is predicted to have molecular function: P-P-bond-hydrolysis-driven protein transmembrane transporter activity. Based on sequence similarity, it is predicted to be involved in the biological process: protein targeting to mitochondrion. 7 alleles are reported. The phenotypes of these alleles are annotated with: trichogen cell; mesothoracic tergum. It has 2 annotated transcripts and 2 annotated polypeptides. Protein features are: Mitochondrial inner membrane translocase complex, Tim8/9/10/13-zinc finger-like. Summary of modENCODE Temporal Expression Profile: Temporal profile ranges from a peak of very high expression to a trough of high expression. Peak expression observed at stages throughout embryogenesis, during early larval stages, during late pupal stages, in stages of adults of both sexes. This gene is annotated by FlyBase as a dicistronic gene, meaning that some or all of its transcripts encode two or more polypeptide-coding open reading frames (ORFs) , with each ORF assigned to a different gene. The distribution of RNA-Seq coverage data amongst the different encoded genes cannot be determined. . |