| Home | Overview of gut regions | Anatomy | Histology | Transgene expression mapping | Gene expression |
| Search expression data by gene: |
| Gene name | inaE | ||||||||||||||||||||||||||||||||||||
| Flybase description | The gene inactivation no afterpotential E is referred to in FlyBase by the symbol Dmel\inaE (CG33174, FBgn0261244). | ||||||||||||||||||||||||||||||||||||
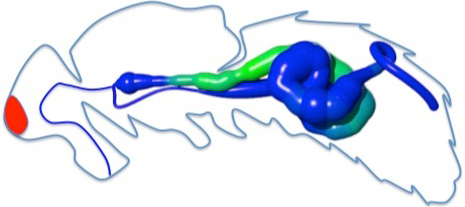
| Expression data along the gut |
|
||||||||||||||||||||||||||||||||||||
| Intestinal gene expression in different physiological conditions | There is not condition-dependent expression data available for this gene. | ||||||||||||||||||||||||||||||||||||
| Gene details (from Flybase) | It is a protein_coding_gene from Drosophila melanogaster. There is experimental evidence that it has the molecular function: lipoprotein lipase activity. There is experimental evidence that it is involved in the biological process: phototransduction. 25 alleles are reported. The phenotypes of these alleles are annotated with: photoreceptor cell R2; photoreceptor cell R1; photoreceptor cell R6; photoreceptor cell R5; photoreceptor cell R4; photoreceptor cell R3. It has 4 annotated transcripts and 4 annotated polypeptides. Protein features are: Lipase, class 3. Summary of modENCODE Temporal Expression Profile: Temporal profile ranges from a peak of moderately high expression to a trough of low expression. Peak expression observed within 00-06 hour embryonic stages. . |